If We Want to Store an Array of Structures Into a Disk File, What File Type Should We Choose?
NumPy: the absolute basics for beginners¶
Welcome to the absolute beginner's guide to NumPy! If you lot take comments or suggestions, please don't hesitate to reach out!
Welcome to NumPy!¶
NumPy (Numerical Python) is an open source Python library that's used in almost every field of scientific discipline and engineering. It's the universal standard for working with numerical information in Python, and it'southward at the core of the scientific Python and PyData ecosystems. NumPy users include anybody from beginning coders to experienced researchers doing state-of-the-art scientific and industrial research and development. The NumPy API is used extensively in Pandas, SciPy, Matplotlib, scikit-acquire, scikit-image and most other data scientific discipline and scientific Python packages.
The NumPy library contains multidimensional array and matrix information structures (yous'll notice more data near this in later sections). Information technology provides ndarray, a homogeneous north-dimensional array object, with methods to efficiently operate on it. NumPy tin exist used to perform a wide diversity of mathematical operations on arrays. It adds powerful data structures to Python that guarantee efficient calculations with arrays and matrices and it supplies an enormous library of high-level mathematical functions that operate on these arrays and matrices.
Learn more than virtually NumPy here!
Installing NumPy¶
To install NumPy, we strongly recommend using a scientific Python distribution. If you're looking for the full instructions for installing NumPy on your operating system, meet Installing NumPy.
If you already have Python, you can install NumPy with:
or
If you don't take Python nonetheless, you might want to consider using Anaconda. It's the easiest way to go started. The skillful thing about getting this distribution is the fact that you don't need to worry too much virtually separately installing NumPy or any of the major packages that you'll be using for your data analyses, like pandas, Scikit-Larn, etc.
How to import NumPy¶
To admission NumPy and its functions import it in your Python code like this:
Nosotros shorten the imported proper noun to np for improve readability of lawmaking using NumPy. This is a widely adopted convention that you should follow so that anyone working with your code tin easily understand it.
Reading the case lawmaking¶
If you aren't already comfortable with reading tutorials that contain a lot of lawmaking, you might not know how to interpret a code block that looks like this:
>>> a = np . arange ( half dozen ) >>> a2 = a [ np . newaxis , :] >>> a2 . shape (ane, 6) If you aren't familiar with this way, it's very easy to sympathize. If you see >>> , you're looking at input, or the code that you would enter. Everything that doesn't accept >>> in front of information technology is output, or the results of running your lawmaking. This is the mode you see when you lot run python on the command line, only if yous're using IPython, you might run into a different manner. Note that it is not part of the lawmaking and will cause an fault if typed or pasted into the Python crush. It tin be safely typed or pasted into the IPython beat out; the >>> is ignored.
What's the divergence between a Python list and a NumPy assortment?¶
NumPy gives y'all an enormous range of fast and efficient means of creating arrays and manipulating numerical data within them. While a Python list tin can contain different data types within a unmarried list, all of the elements in a NumPy array should be homogeneous. The mathematical operations that are meant to exist performed on arrays would be extremely inefficient if the arrays weren't homogeneous.
Why utilise NumPy?
NumPy arrays are faster and more compact than Python lists. An assortment consumes less memory and is convenient to employ. NumPy uses much less memory to store data and it provides a mechanism of specifying the data types. This allows the code to exist optimized fifty-fifty further.
What is an array?¶
An array is a key data structure of the NumPy library. An array is a grid of values and it contains information about the raw information, how to locate an chemical element, and how to interpret an chemical element. It has a filigree of elements that can be indexed in various ways. The elements are all of the same type, referred to as the assortment dtype .
An assortment can be indexed past a tuple of nonnegative integers, by booleans, by another array, or by integers. The rank of the array is the number of dimensions. The shape of the array is a tuple of integers giving the size of the array along each dimension.
I way we can initialize NumPy arrays is from Python lists, using nested lists for two- or higher-dimensional data.
For example:
>>> a = np . array ([ i , 2 , iii , 4 , 5 , 6 ]) or:
>>> a = np . array ([[ one , 2 , 3 , 4 ], [ 5 , 6 , vii , eight ], [ 9 , x , 11 , 12 ]]) Nosotros tin can access the elements in the array using square brackets. When you lot're accessing elements, remember that indexing in NumPy starts at 0. That means that if you want to admission the first element in your array, you'll be accessing element "0".
>>> print ( a [ 0 ]) [1 2 3 4] More data about arrays¶
This section covers 1D array , 2D array , ndarray , vector , matrix
You might occasionally hear an array referred to as a "ndarray," which is shorthand for "Northward-dimensional array." An N-dimensional array is simply an array with any number of dimensions. You might likewise hear 1-D, or one-dimensional assortment, ii-D, or two-dimensional array, then on. The NumPy ndarray grade is used to stand for both matrices and vectors. A vector is an array with a single dimension (there's no difference between row and column vectors), while a matrix refers to an array with two dimensions. For three-D or higher dimensional arrays, the term tensor is also commonly used.
What are the attributes of an assortment?
An array is commonly a fixed-size container of items of the same type and size. The number of dimensions and items in an array is divers by its shape. The shape of an array is a tuple of non-negative integers that specify the sizes of each dimension.
In NumPy, dimensions are called axes. This means that if yous have a 2nd array that looks like this:
[[ 0. , 0. , 0. ], [ 1. , 1. , 1. ]] Your array has 2 axes. The outset axis has a length of 2 and the second centrality has a length of 3.
But like in other Python container objects, the contents of an array can be accessed and modified by indexing or slicing the array. Unlike the typical container objects, dissimilar arrays can share the same data, so changes made on one array might be visible in some other.
Array attributes reflect information intrinsic to the array itself. If you demand to go, or even fix, backdrop of an assortment without creating a new array, you lot tin can ofttimes access an array through its attributes.
Read more about array attributes here and learn virtually assortment objects here.
How to create a bones array¶
This section covers np.array() , np.zeros() , np.ones() , np.empty() , np.arange() , np.linspace() , dtype
To create a NumPy array, you can use the function np.array() .
All you need to do to create a unproblematic assortment is pass a list to information technology. If you lot choose to, y'all can also specify the type of information in your listing. You tin find more information about data types here.
>>> import numpy equally np >>> a = np . array ([ ane , 2 , 3 ]) You can visualize your assortment this way:

Exist aware that these visualizations are meant to simplify ideas and give you a basic understanding of NumPy concepts and mechanics. Arrays and array operations are much more complicated than are captured hither!
As well creating an assortment from a sequence of elements, you can hands create an array filled with 0 's:
>>> np . zeros ( 2 ) assortment([0., 0.]) Or an array filled with 1 's:
>>> np . ones ( 2 ) array([1., 1.]) Or even an empty array! The office empty creates an assortment whose initial content is random and depends on the state of the memory. The reason to utilise empty over zeros (or something like) is speed - merely brand sure to fill every element afterwards!
>>> # Create an empty array with 2 elements >>> np . empty ( 2 ) assortment([ three.14, 42. ]) # may vary You tin create an array with a range of elements:
>>> np . arange ( 4 ) array([0, 1, 2, 3]) And even an array that contains a range of evenly spaced intervals. To exercise this, you will specify the showtime number, last number, and the step size.
>>> np . arange ( 2 , nine , 2 ) array([2, 4, 6, 8]) Y'all can also use np.linspace() to create an array with values that are spaced linearly in a specified interval:
>>> np . linspace ( 0 , 10 , num = 5 ) assortment([ 0. , ii.five, 5. , 7.five, 10. ]) Specifying your data type
While the default data type is floating point ( np.float64 ), y'all tin can explicitly specify which data type you want using the dtype keyword.
>>> x = np . ones ( 2 , dtype = np . int64 ) >>> x array([1, one]) Acquire more about creating arrays here
Calculation, removing, and sorting elements¶
This section covers np.sort() , np.concatenate()
Sorting an element is unproblematic with np.sort() . You can specify the centrality, kind, and order when you call the office.
If you start with this array:
>>> arr = np . array ([ 2 , 1 , 5 , 3 , seven , 4 , vi , 8 ]) You can quickly sort the numbers in ascending order with:
>>> np . sort ( arr ) array([1, two, 3, four, 5, half-dozen, 7, 8]) In improver to sort, which returns a sorted copy of an array, y'all can use:
-
argsort, which is an indirect sort along a specified centrality, -
lexsort, which is an indirect stable sort on multiple keys, -
searchsorted, which will find elements in a sorted array, and -
sectionalisation, which is a partial sort.
To read more about sorting an array, see: sort .
If you start with these arrays:
>>> a = np . array ([ 1 , 2 , 3 , 4 ]) >>> b = np . array ([ 5 , 6 , 7 , viii ]) You lot tin can concatenate them with np.concatenate() .
>>> np . concatenate (( a , b )) array([i, 2, three, iv, 5, 6, 7, eight]) Or, if you lot start with these arrays:
>>> x = np . array ([[ one , 2 ], [ 3 , four ]]) >>> y = np . assortment ([[ 5 , 6 ]]) You lot can concatenate them with:
>>> np . concatenate (( ten , y ), axis = 0 ) array([[one, 2], [3, 4], [5, six]]) In social club to remove elements from an array, information technology's simple to use indexing to select the elements that you want to go along.
To read more about concatenate, come across: concatenate .
How do you know the shape and size of an array?¶
This department covers ndarray.ndim , ndarray.size , ndarray.shape
ndarray.ndim volition tell you the number of axes, or dimensions, of the assortment.
ndarray.size volition tell you the total number of elements of the assortment. This is the product of the elements of the array's shape.
ndarray.shape will display a tuple of integers that indicate the number of elements stored along each dimension of the assortment. If, for example, you have a ii-D array with 2 rows and 3 columns, the shape of your array is (two, 3) .
For instance, if you create this array:
>>> array_example = np . array ([[[ 0 , 1 , 2 , 3 ], ... [ four , five , 6 , 7 ]], ... ... [[ 0 , i , 2 , 3 ], ... [ four , v , 6 , vii ]], ... ... [[ 0 , i , ii , three ], ... [ 4 , 5 , half dozen , 7 ]]]) To find the number of dimensions of the array, run:
To detect the full number of elements in the array, run:
>>> array_example . size 24 And to notice the shape of your array, run:
>>> array_example . shape (3, 2, 4) Can yous reshape an array?¶
This department covers arr.reshape()
Yes!
Using arr.reshape() will give a new shape to an array without irresolute the data. Just remember that when you use the reshape method, the assortment y'all want to produce needs to have the same number of elements as the original assortment. If you lot start with an array with 12 elements, yous'll demand to make sure that your new array also has a total of 12 elements.
If you start with this array:
>>> a = np . arange ( 6 ) >>> print ( a ) [0 1 2 3 iv 5] You can use reshape() to reshape your array. For example, you can reshape this assortment to an array with three rows and two columns:
>>> b = a . reshape ( 3 , two ) >>> print ( b ) [[0 1] [2 three] [4 v]] With np.reshape , y'all tin specify a few optional parameters:
>>> np . reshape ( a , newshape = ( i , half-dozen ), social club = 'C' ) assortment([[0, ane, two, 3, iv, v]]) a is the array to be reshaped.
newshape is the new shape you want. Yous tin can specify an integer or a tuple of integers. If you lot specify an integer, the result will be an assortment of that length. The shape should be compatible with the original shape.
order: C means to read/write the elements using C-like index society, F means to read/write the elements using Fortran-like index order, A ways to read/write the elements in Fortran-like index gild if a is Fortran face-to-face in retentivity, C-similar order otherwise. (This is an optional parameter and doesn't need to be specified.)
If you desire to acquire more than about C and Fortran order, you can read more about the internal organization of NumPy arrays hither. Substantially, C and Fortran orders have to practice with how indices stand for to the order the array is stored in memory. In Fortran, when moving through the elements of a two-dimensional array as information technology is stored in retentivity, the first index is the about rapidly varying index. As the beginning index moves to the side by side row as it changes, the matrix is stored ane column at a time. This is why Fortran is idea of every bit a Column-major language. In C on the other mitt, the final index changes the most chop-chop. The matrix is stored by rows, making it a Row-major language. What y'all do for C or Fortran depends on whether it's more than important to preserve the indexing convention or not reorder the data.
Learn more about shape manipulation hither.
How to convert a 1D array into a 2d array (how to add together a new axis to an assortment)¶
This department covers np.newaxis , np.expand_dims
You can use np.newaxis and np.expand_dims to increment the dimensions of your existing array.
Using np.newaxis will increment the dimensions of your array by 1 dimension when used once. This means that a 1D array will become a 2D array, a 2D array will go a 3D array, and then on.
For case, if you starting time with this array:
>>> a = np . assortment ([ 1 , 2 , 3 , 4 , 5 , 6 ]) >>> a . shape (6,) Yous can use np.newaxis to add a new axis:
>>> a2 = a [ np . newaxis , :] >>> a2 . shape (1, 6) Yous tin explicitly convert a 1D array with either a row vector or a column vector using np.newaxis . For instance, you can convert a 1D array to a row vector by inserting an axis along the starting time dimension:
>>> row_vector = a [ np . newaxis , :] >>> row_vector . shape (ane, vi) Or, for a cavalcade vector, y'all tin insert an centrality along the second dimension:
>>> col_vector = a [:, np . newaxis ] >>> col_vector . shape (half dozen, one) You can also expand an array by inserting a new axis at a specified position with np.expand_dims .
For example, if y'all outset with this array:
>>> a = np . array ([ i , 2 , 3 , 4 , v , 6 ]) >>> a . shape (half-dozen,) You tin can employ np.expand_dims to add an axis at index position i with:
>>> b = np . expand_dims ( a , centrality = ane ) >>> b . shape (six, one) You tin can add together an axis at alphabetize position 0 with:
>>> c = np . expand_dims ( a , axis = 0 ) >>> c . shape (1, 6) Find more than information about newaxis here and expand_dims at expand_dims .
Indexing and slicing¶
Yous can index and slice NumPy arrays in the same ways you lot can slice Python lists.
>>> data = np . assortment ([ i , 2 , 3 ]) >>> information [ i ] 2 >>> data [ 0 : 2 ] assortment([1, 2]) >>> information [ i :] array([2, 3]) >>> data [ - ii :] array([2, iii]) You can visualize it this way:
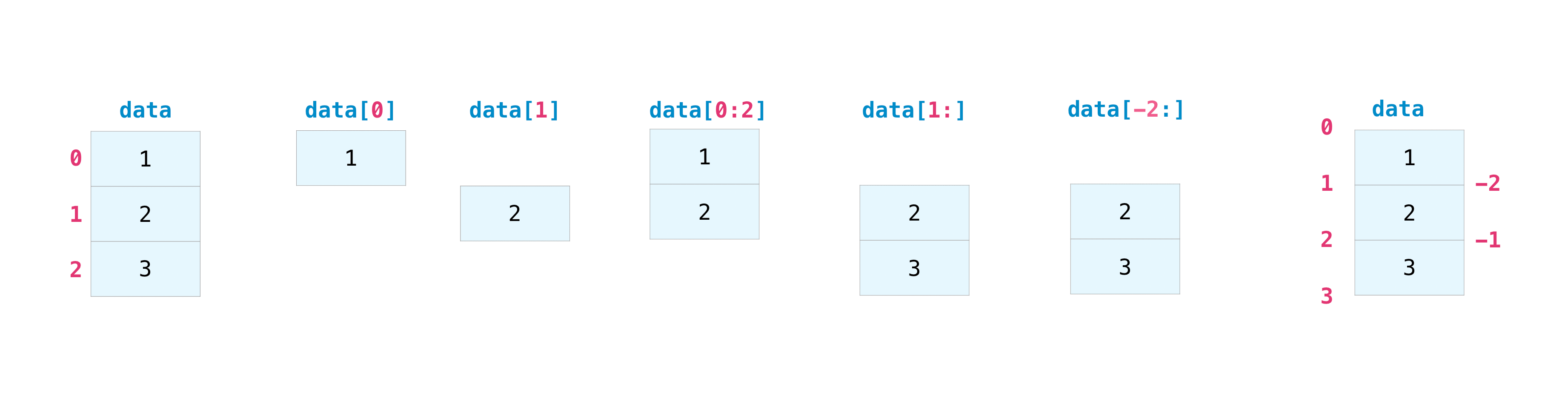

Yous may desire to take a department of your array or specific array elements to utilize in further analysis or boosted operations. To do that, y'all'll need to subset, piece, and/or index your arrays.
If y'all want to select values from your array that fulfill sure conditions, information technology'due south straightforward with NumPy.
For example, if you commencement with this array:
>>> a = np . array ([[ 1 , ii , 3 , iv ], [ 5 , 6 , seven , 8 ], [ 9 , 10 , 11 , 12 ]]) Yous tin hands impress all of the values in the array that are less than 5.
>>> print ( a [ a < 5 ]) [1 2 3 4] You can likewise select, for case, numbers that are equal to or greater than 5, and use that condition to index an assortment.
>>> five_up = ( a >= 5 ) >>> print ( a [ five_up ]) [ five six vii 8 nine 10 11 12] You can select elements that are divisible by 2:
>>> divisible_by_2 = a [ a % 2 == 0 ] >>> print ( divisible_by_2 ) [ 2 4 6 eight 10 12] Or you can select elements that satisfy two weather condition using the & and | operators:
>>> c = a [( a > ii ) & ( a < eleven )] >>> impress ( c ) [ 3 four v 6 seven 8 ix 10] You can also make use of the logical operators & and | in order to render boolean values that specify whether or non the values in an assortment fulfill a sure condition. This tin be useful with arrays that contain names or other chiselled values.
>>> five_up = ( a > five ) | ( a == 5 ) >>> impress ( five_up ) [[False False False Faux] [ True Truthful True True] [ True True True Truthful]] You lot tin can too use np.nonzero() to select elements or indices from an array.
Starting with this array:
>>> a = np . array ([[ 1 , 2 , 3 , 4 ], [ 5 , 6 , 7 , 8 ], [ ix , x , xi , 12 ]]) You can use np.nonzero() to impress the indices of elements that are, for instance, less than 5:
>>> b = np . nonzero ( a < v ) >>> print ( b ) (array([0, 0, 0, 0]), array([0, 1, 2, 3])) In this instance, a tuple of arrays was returned: ane for each dimension. The first array represents the row indices where these values are found, and the 2d assortment represents the column indices where the values are found.
If you want to generate a listing of coordinates where the elements exist, you can zip the arrays, iterate over the listing of coordinates, and impress them. For example:
>>> list_of_coordinates = list ( nothing ( b [ 0 ], b [ ane ])) >>> for coord in list_of_coordinates : ... print ( coord ) (0, 0) (0, 1) (0, 2) (0, 3) You can besides use np.nonzero() to print the elements in an array that are less than v with:
>>> print ( a [ b ]) [1 2 3 4] If the element you're looking for doesn't exist in the array, then the returned assortment of indices volition be empty. For case:
>>> not_there = np . nonzero ( a == 42 ) >>> print ( not_there ) (array([], dtype=int64), array([], dtype=int64)) Learn more about indexing and slicing here and here.
Read more about using the nonzero part at: nonzero .
How to create an array from existing information¶
This department covers slicing and indexing , np.vstack() , np.hstack() , np.hsplit() , .view() , re-create()
You can easily create a new array from a section of an existing array.
Let's say y'all have this array:
>>> a = np . assortment ([ 1 , ii , 3 , 4 , v , 6 , 7 , 8 , ix , 10 ]) You can create a new array from a department of your array any time past specifying where you want to slice your array.
>>> arr1 = a [ iii : viii ] >>> arr1 array([4, 5, 6, 7, 8]) Hither, you lot grabbed a section of your array from index position 3 through index position 8.
You can also stack two existing arrays, both vertically and horizontally. Allow's say you have ii arrays, a1 and a2 :
>>> a1 = np . array ([[ i , i ], ... [ ii , two ]]) >>> a2 = np . array ([[ 3 , 3 ], ... [ iv , 4 ]]) Yous can stack them vertically with vstack :
>>> np . vstack (( a1 , a2 )) array([[1, ane], [2, ii], [3, iii], [4, 4]]) Or stack them horizontally with hstack :
>>> np . hstack (( a1 , a2 )) assortment([[i, one, three, 3], [2, 2, 4, 4]]) Y'all can separate an array into several smaller arrays using hsplit . You can specify either the number of every bit shaped arrays to return or the columns after which the division should occur.
Permit'south say you have this array:
>>> x = np . arange ( i , 25 ) . reshape ( 2 , 12 ) >>> x assortment([[ 1, 2, three, 4, 5, half dozen, 7, eight, 9, 10, 11, 12], [thirteen, 14, 15, xvi, 17, xviii, 19, twenty, 21, 22, 23, 24]]) If you wanted to split this array into three equally shaped arrays, you would run:
>>> np . hsplit ( x , 3 ) [array([[1, 2, 3, 4], [thirteen, 14, 15, sixteen]]), assortment([[ 5, vi, 7, eight], [17, 18, 19, xx]]), array([[ 9, 10, 11, 12], [21, 22, 23, 24]])] If you wanted to carve up your assortment subsequently the third and 4th column, you lot'd run:
>>> np . hsplit ( x , ( three , 4 )) [array([[i, two, 3], [13, xiv, fifteen]]), array([[ 4], [16]]), array([[ 5, half dozen, 7, 8, ix, 10, xi, 12], [17, 18, 19, 20, 21, 22, 23, 24]])] Learn more than virtually stacking and splitting arrays here.
Yous can use the view method to create a new assortment object that looks at the aforementioned data as the original array (a shallow re-create).
Views are an of import NumPy concept! NumPy functions, every bit well as operations like indexing and slicing, volition return views whenever possible. This saves memory and is faster (no copy of the data has to exist made). However it'south important to be aware of this - modifying data in a view also modifies the original array!
Permit'south say you create this array:
>>> a = np . array ([[ 1 , 2 , three , 4 ], [ 5 , half-dozen , 7 , eight ], [ nine , ten , 11 , 12 ]]) Now we create an array b1 by slicing a and modify the first element of b1 . This will modify the corresponding element in a as well!
>>> b1 = a [ 0 , :] >>> b1 array([1, 2, iii, 4]) >>> b1 [ 0 ] = 99 >>> b1 array([99, 2, 3, 4]) >>> a array([[99, ii, three, four], [ five, 6, 7, 8], [ 9, x, eleven, 12]]) Using the copy method volition make a complete copy of the array and its data (a deep copy). To utilise this on your assortment, you lot could run:
Learn more about copies and views here.
Basic array operations¶
This department covers add-on, subtraction, multiplication, division, and more than
One time you've created your arrays, you can start to work with them. Let'due south say, for example, that you've created two arrays, one chosen "data" and i called "ones"
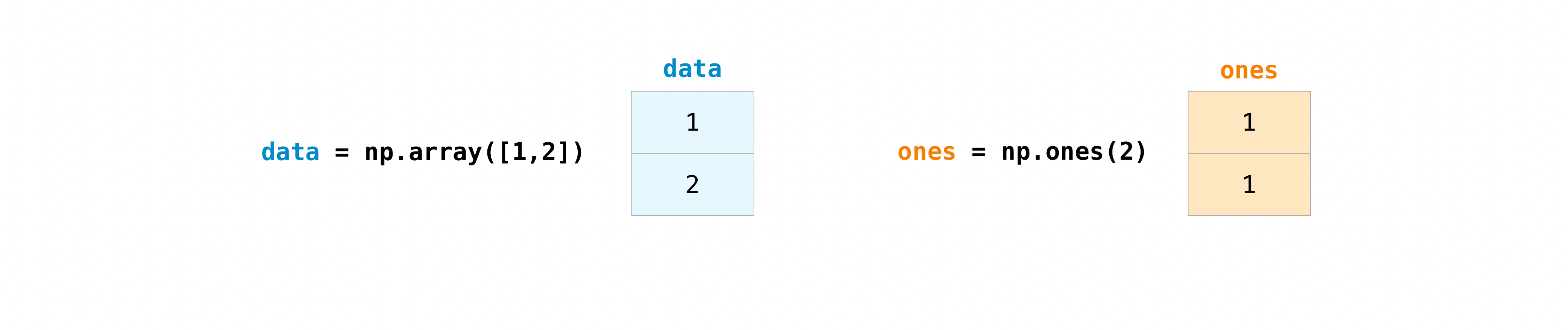
You can add the arrays together with the plus sign.
>>> data = np . array ([ 1 , 2 ]) >>> ones = np . ones ( 2 , dtype = int ) >>> data + ones array([2, 3]) 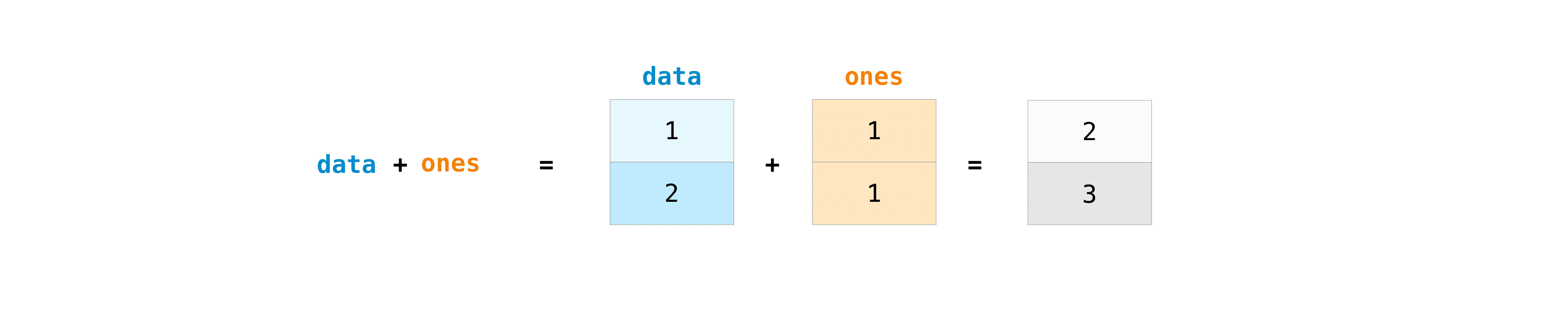
You tin can, of course, practise more than just add-on!
>>> data - ones array([0, 1]) >>> data * data array([one, 4]) >>> data / data array([i., 1.]) 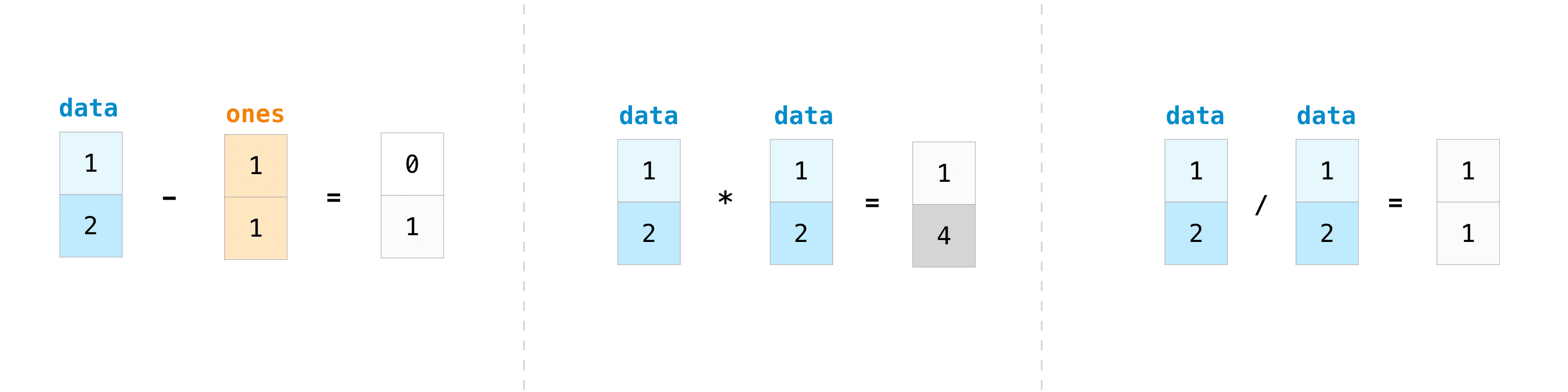
Basic operations are uncomplicated with NumPy. If you want to notice the sum of the elements in an assortment, y'all'd use sum() . This works for 1D arrays, 2D arrays, and arrays in higher dimensions.
>>> a = np . array ([ 1 , 2 , 3 , 4 ]) >>> a . sum () x To add the rows or the columns in a second array, you would specify the centrality.
If you lot start with this array:
>>> b = np . array ([[ 1 , ane ], [ ii , 2 ]]) You lot can sum over the axis of rows with:
>>> b . sum ( axis = 0 ) array([3, 3]) You can sum over the axis of columns with:
>>> b . sum ( axis = 1 ) array([two, iv]) Learn more almost basic operations here.
Broadcasting¶
There are times when you might want to carry out an operation between an array and a single number (also called an operation betwixt a vector and a scalar) or betwixt arrays of ii unlike sizes. For example, your array (we'll call it "data") might contain data about distance in miles but you want to catechumen the information to kilometers. You can perform this functioning with:
>>> data = np . array ([ 1.0 , 2.0 ]) >>> information * 1.vi array([one.half dozen, 3.2]) 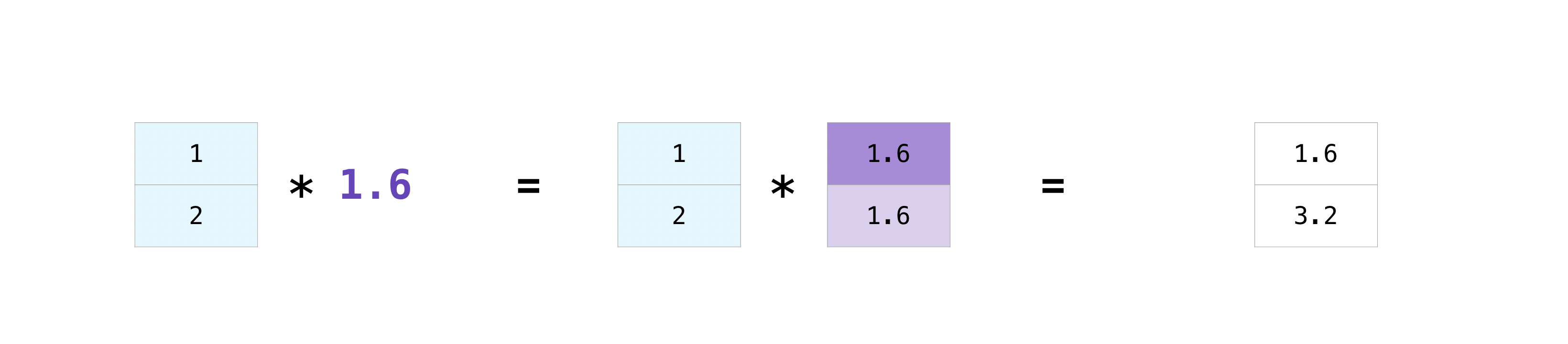
NumPy understands that the multiplication should happen with each cell. That concept is called broadcasting. Broadcasting is a mechanism that allows NumPy to perform operations on arrays of unlike shapes. The dimensions of your assortment must exist compatible, for instance, when the dimensions of both arrays are equal or when one of them is 1. If the dimensions are not uniform, you will get a ValueError .
Learn more about broadcasting here.
More useful array operations¶
This section covers maximum, minimum, sum, mean, product, standard divergence, and more than
NumPy too performs aggregation functions. In add-on to min , max , and sum , yous can easily run hateful to get the average, prod to become the upshot of multiplying the elements together, std to get the standard divergence, and more.
>>> data . max () ii.0 >>> data . min () i.0 >>> data . sum () 3.0 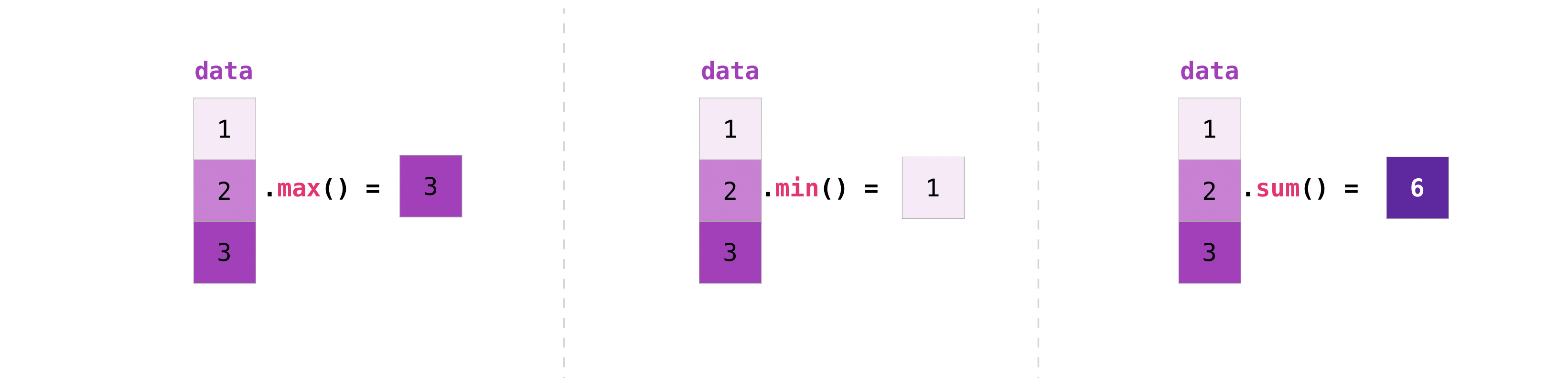
Permit'due south kickoff with this array, called "a"
>>> a = np . array ([[ 0.45053314 , 0.17296777 , 0.34376245 , 0.5510652 ], ... [ 0.54627315 , 0.05093587 , 0.40067661 , 0.55645993 ], ... [ 0.12697628 , 0.82485143 , 0.26590556 , 0.56917101 ]]) It's very common to want to aggregate forth a row or cavalcade. Past default, every NumPy aggregation function will return the aggregate of the entire assortment. To find the sum or the minimum of the elements in your assortment, run:
Or:
You lot can specify on which axis you want the assemblage function to be computed. For example, y'all tin can notice the minimum value inside each column by specifying axis=0 .
>>> a . min ( axis = 0 ) array([0.12697628, 0.05093587, 0.26590556, 0.5510652 ]) The 4 values listed higher up correspond to the number of columns in your array. With a four-column array, you lot will get four values as your result.
Read more about array methods here.
Creating matrices¶
You can pass Python lists of lists to create a ii-D array (or "matrix") to represent them in NumPy.
>>> information = np . array ([[ 1 , ii ], [ 3 , 4 ], [ v , half dozen ]]) >>> information assortment([[one, 2], [three, 4], [5, 6]])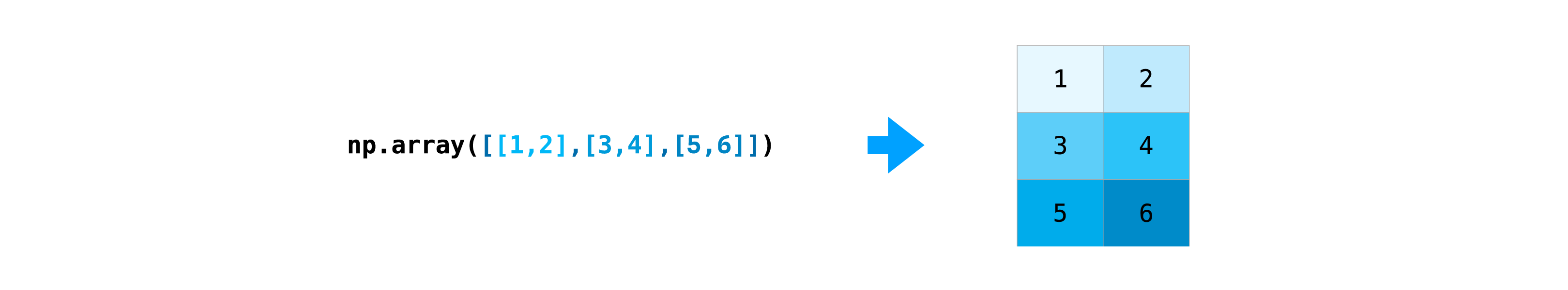 
Indexing and slicing operations are useful when you're manipulating matrices:
>>> data [ 0 , i ] 2 >>> data [ one : 3 ] array([[three, iv], [5, 6]]) >>> data [ 0 : 2 , 0 ] assortment([1, three])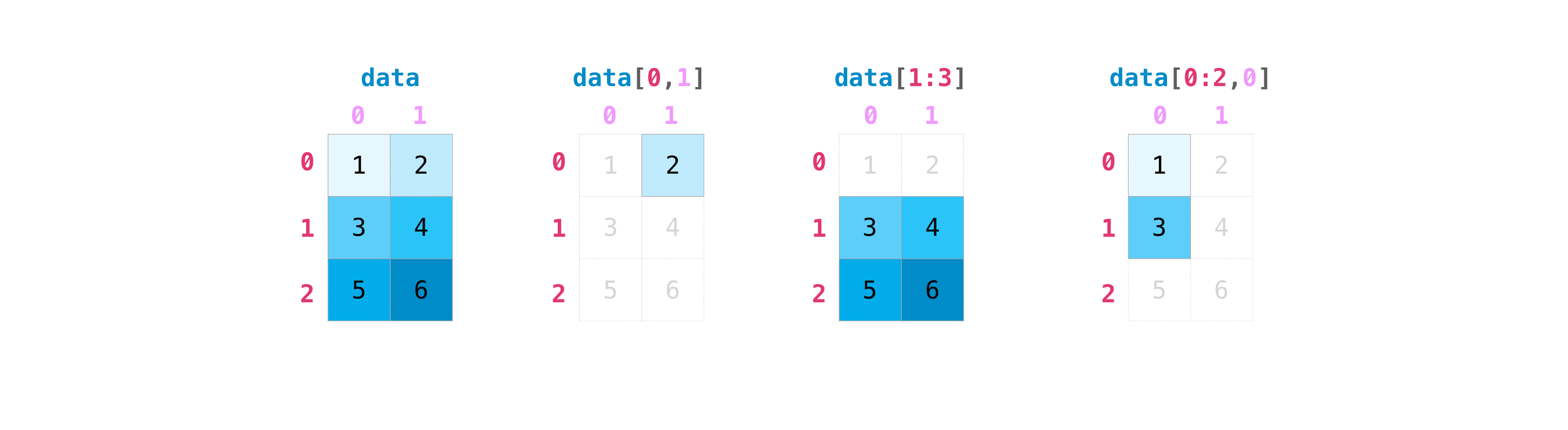 
You lot tin can amass matrices the same mode you aggregated vectors:
>>> data . max () 6 >>> information . min () 1 >>> information . sum () 21 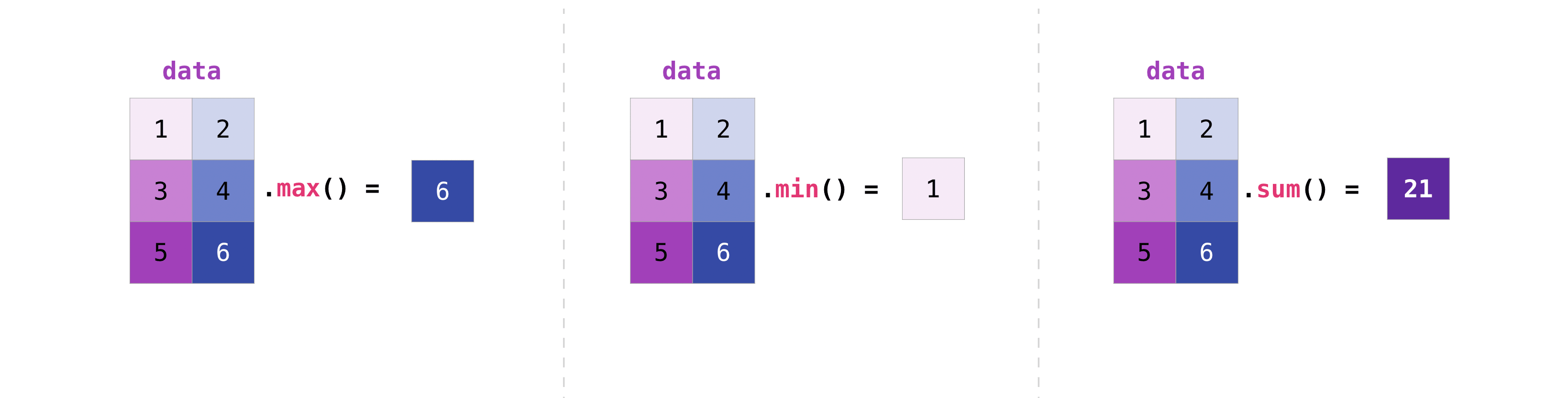
You tin can amass all the values in a matrix and you lot tin aggregate them across columns or rows using the centrality parameter. To illustrate this point, let'southward wait at a slightly modified dataset:
>>> data = np . assortment ([[ 1 , 2 ], [ 5 , 3 ], [ 4 , vi ]]) >>> information array([[1, 2], [v, 3], [four, six]]) >>> information . max ( axis = 0 ) array([five, 6]) >>> information . max ( axis = 1 ) assortment([ii, 5, 6]) 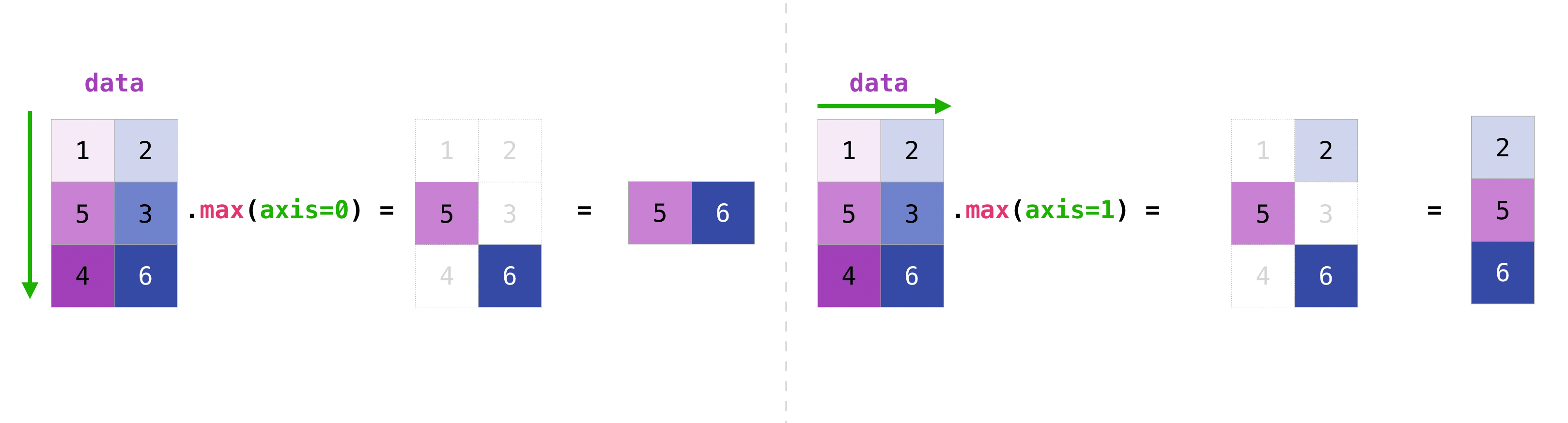
Once you've created your matrices, you tin add together and multiply them using arithmetic operators if you have ii matrices that are the aforementioned size.
>>> data = np . array ([[ one , 2 ], [ 3 , 4 ]]) >>> ones = np . assortment ([[ i , 1 ], [ ane , 1 ]]) >>> data + ones array([[2, 3], [four, v]]) 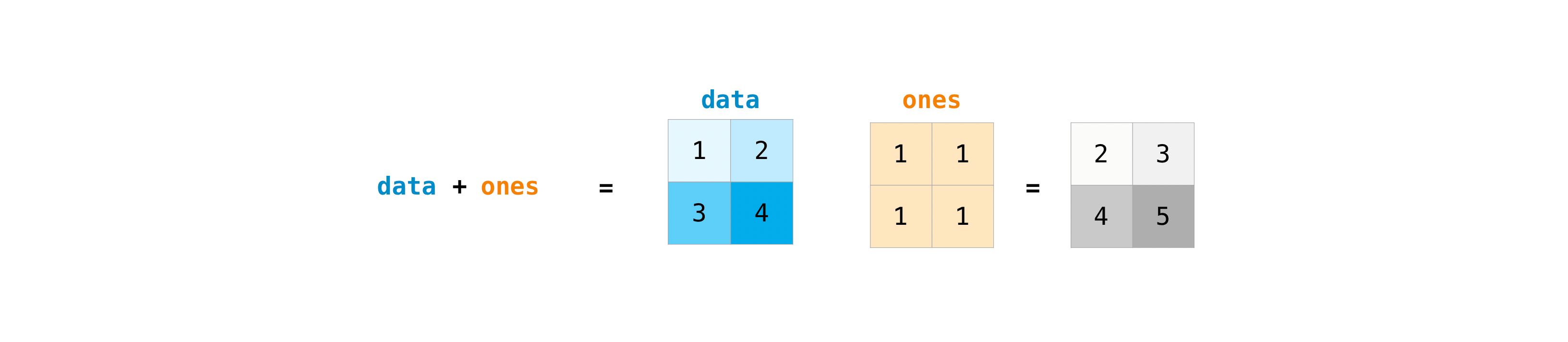
Y'all can exercise these arithmetic operations on matrices of different sizes, but only if one matrix has only one column or one row. In this case, NumPy will use its broadcast rules for the operation.
>>> data = np . array ([[ 1 , 2 ], [ 3 , 4 ], [ 5 , 6 ]]) >>> ones_row = np . array ([[ 1 , 1 ]]) >>> data + ones_row array([[2, 3], [4, v], [6, 7]]) 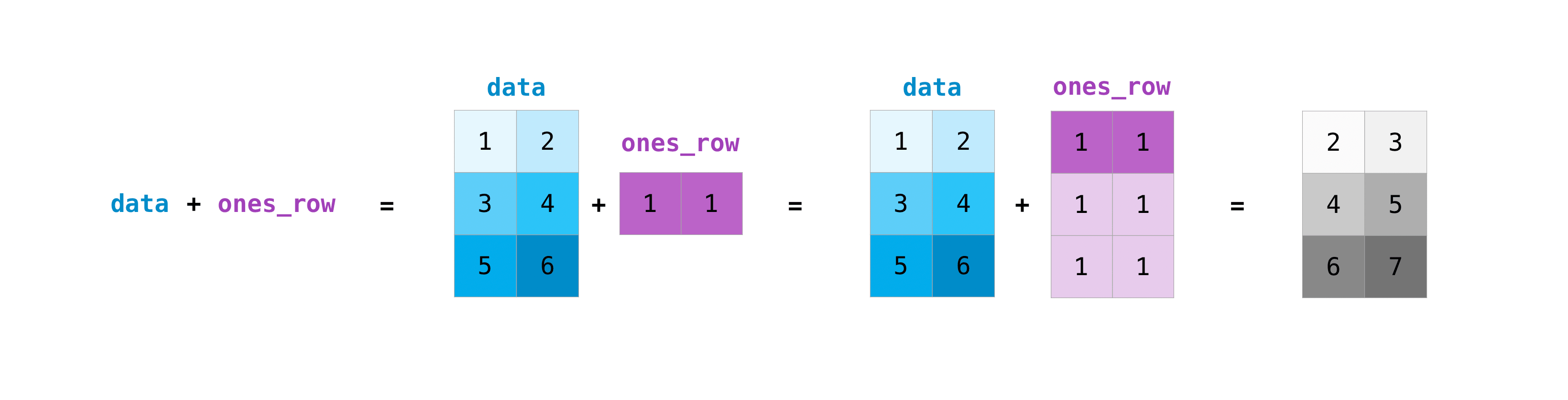
Be aware that when NumPy prints Northward-dimensional arrays, the concluding axis is looped over the fastest while the first centrality is the slowest. For case:
>>> np . ones (( iv , 3 , 2 )) assortment([[[1., 1.], [1., 1.], [1., ane.]], [[1., i.], [i., i.], [one., 1.]], [[one., 1.], [i., one.], [1., one.]], [[1., 1.], [1., 1.], [1., 1.]]]) In that location are frequently instances where we desire NumPy to initialize the values of an array. NumPy offers functions like ones() and zeros() , and the random.Generator class for random number generation for that. All yous need to do is pass in the number of elements you want information technology to generate:
>>> np . ones ( 3 ) array([one., 1., one.]) >>> np . zeros ( 3 ) assortment([0., 0., 0.]) # the simplest style to generate random numbers >>> rng = np . random . default_rng ( 0 ) >>> rng . random ( 3 ) array([0.63696169, 0.26978671, 0.04097352]) 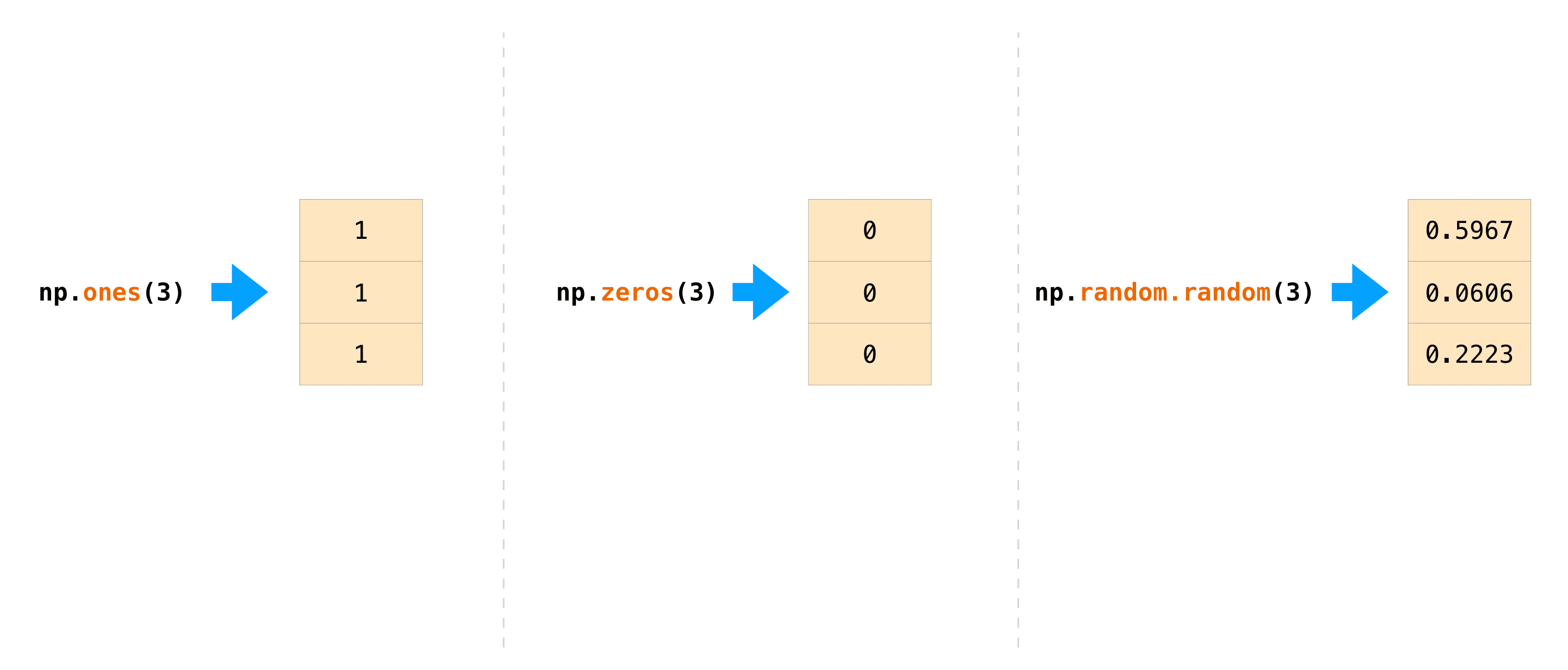
You can also use ones() , zeros() , and random() to create a second array if you requite them a tuple describing the dimensions of the matrix:
>>> np . ones (( 3 , two )) array([[1., 1.], [1., i.], [1., ane.]]) >>> np . zeros (( iii , two )) assortment([[0., 0.], [0., 0.], [0., 0.]]) >>> rng . random (( 3 , two )) array([[0.01652764, 0.81327024], [0.91275558, 0.60663578], [0.72949656, 0.54362499]]) # may vary 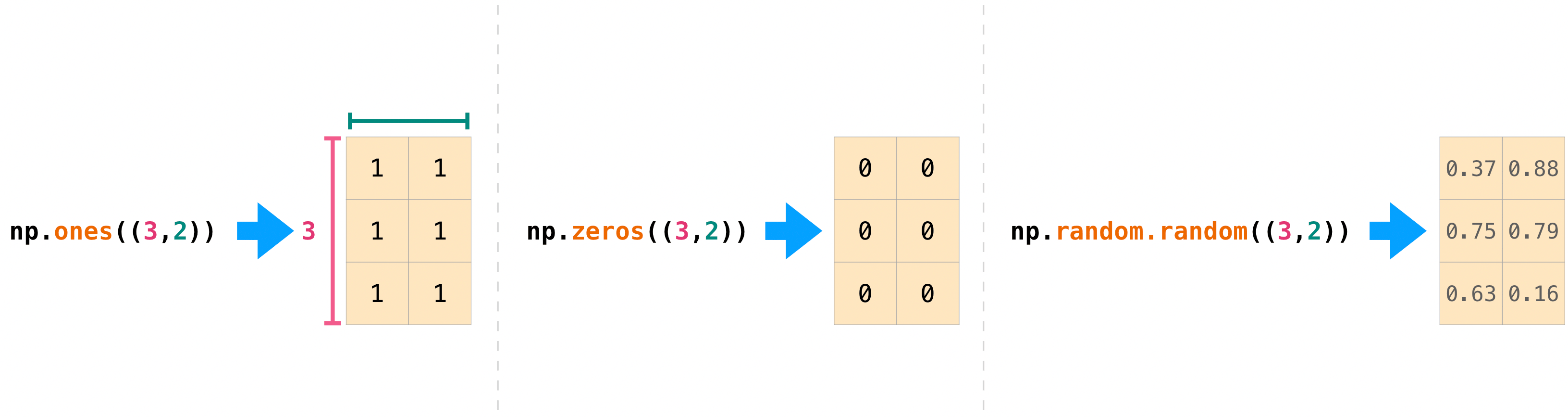
Read more about creating arrays, filled with 0 'south, 1 's, other values or uninitialized, at array creation routines.
Generating random numbers¶
The use of random number generation is an important function of the configuration and evaluation of many numerical and machine learning algorithms. Whether you lot need to randomly initialize weights in an artificial neural network, split data into random sets, or randomly shuffle your dataset, existence able to generate random numbers (actually, repeatable pseudo-random numbers) is essential.
With Generator.integers , you tin can generate random integers from low (recollect that this is inclusive with NumPy) to high (exclusive). You can set endpoint=Truthful to brand the high number inclusive.
You can generate a 2 x iv array of random integers between 0 and 4 with:
>>> rng . integers ( 5 , size = ( ii , iv )) array([[2, one, 1, 0], [0, 0, 0, 4]]) # may vary Read more about random number generation here.
How to get unique items and counts¶
This section covers np.unique()
Y'all tin can discover the unique elements in an array easily with np.unique .
For case, if you start with this array:
>>> a = np . array ([ 11 , xi , 12 , 13 , fourteen , xv , sixteen , 17 , 12 , xiii , xi , 14 , eighteen , 19 , 20 ]) you can utilize np.unique to print the unique values in your array:
>>> unique_values = np . unique ( a ) >>> impress ( unique_values ) [11 12 thirteen 14 fifteen 16 17 eighteen 19 20] To get the indices of unique values in a NumPy array (an array of first alphabetize positions of unique values in the assortment), just pass the return_index statement in np.unique() as well as your array.
>>> unique_values , indices_list = np . unique ( a , return_index = True ) >>> impress ( indices_list ) [ 0 ii 3 iv 5 6 7 12 13 14] Yous can pass the return_counts argument in np.unique() along with your array to go the frequency count of unique values in a NumPy array.
>>> unique_values , occurrence_count = np . unique ( a , return_counts = Truthful ) >>> print ( occurrence_count ) [iii 2 2 2 1 i 1 i 1 1] This also works with 2d arrays! If y'all beginning with this array:
>>> a_2d = np . assortment ([[ 1 , 2 , three , 4 ], [ 5 , six , 7 , 8 ], [ 9 , ten , xi , 12 ], [ 1 , two , iii , 4 ]]) You can discover unique values with:
>>> unique_values = np . unique ( a_2d ) >>> impress ( unique_values ) [ 1 2 3 4 5 six 7 eight nine ten eleven 12] If the axis argument isn't passed, your 2d assortment will be flattened.
If you lot desire to go the unique rows or columns, brand sure to laissez passer the axis argument. To find the unique rows, specify axis=0 and for columns, specify axis=one .
>>> unique_rows = np . unique ( a_2d , axis = 0 ) >>> impress ( unique_rows ) [[ 1 2 iii four] [ 5 6 7 8] [ ix 10 11 12]] To get the unique rows, index position, and occurrence count, you can apply:
>>> unique_rows , indices , occurrence_count = np . unique ( ... a_2d , axis = 0 , return_counts = Truthful , return_index = True ) >>> print ( unique_rows ) [[ ane two three four] [ 5 6 7 eight] [ nine 10 11 12]] >>> impress ( indices ) [0 ane 2] >>> print ( occurrence_count ) [ii 1 1] To learn more about finding the unique elements in an array, meet unique .
Transposing and reshaping a matrix¶
This section covers arr.reshape() , arr.transpose() , arr.T
It's mutual to need to transpose your matrices. NumPy arrays accept the property T that allows y'all to transpose a matrix.
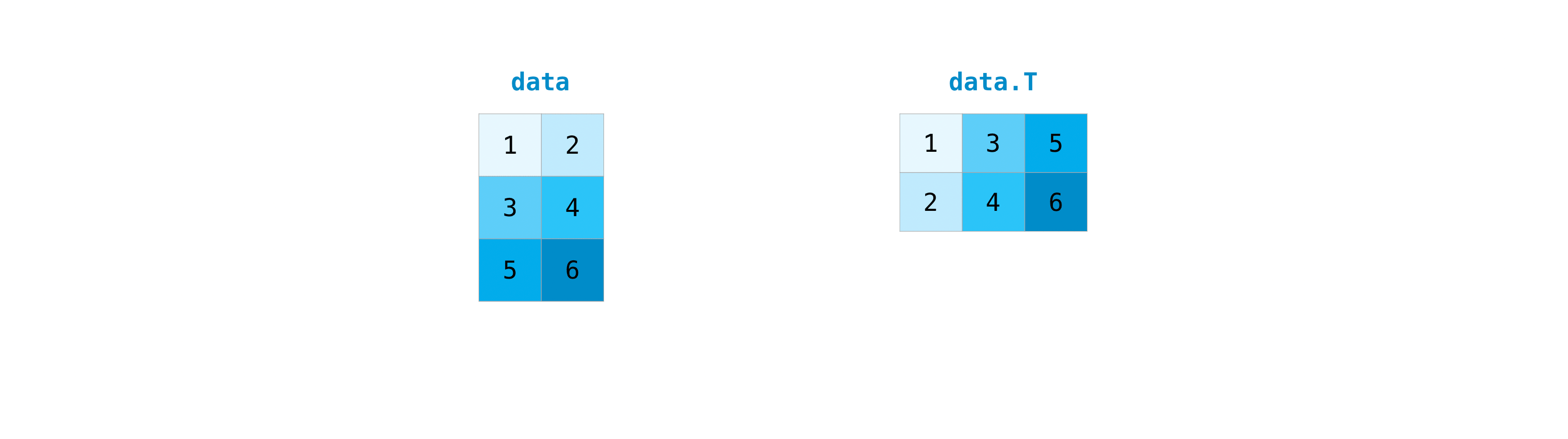
You lot may also need to switch the dimensions of a matrix. This can happen when, for example, yous have a model that expects a certain input shape that is different from your dataset. This is where the reshape method tin can be useful. You simply need to laissez passer in the new dimensions that y'all want for the matrix.
>>> information . reshape ( two , 3 ) array([[1, 2, 3], [4, v, half dozen]]) >>> data . reshape ( 3 , 2 ) array([[1, 2], [3, 4], [v, half-dozen]]) 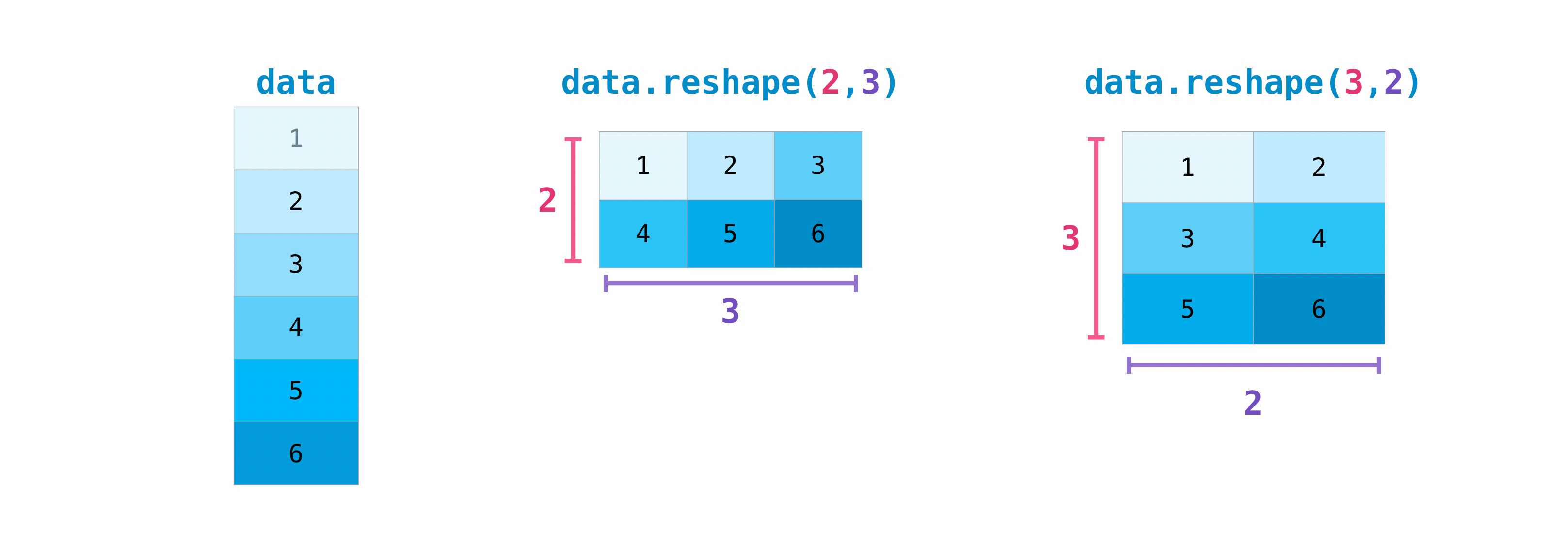
Yous tin too use .transpose() to contrary or alter the axes of an array co-ordinate to the values yous specify.
If you starting time with this array:
>>> arr = np . arange ( 6 ) . reshape (( 2 , 3 )) >>> arr array([[0, 1, 2], [three, four, 5]]) You can transpose your array with arr.transpose() .
>>> arr . transpose () assortment([[0, three], [1, iv], [two, v]]) You tin can also utilize arr.T :
>>> arr . T array([[0, three], [ane, 4], [2, v]]) To learn more than about transposing and reshaping arrays, run across transpose and reshape .
How to contrary an array¶
This section covers np.flip()
NumPy's np.flip() office allows you to flip, or reverse, the contents of an array along an axis. When using np.flip() , specify the array yous would similar to opposite and the axis. If you don't specify the axis, NumPy will reverse the contents along all of the axes of your input assortment.
Reversing a 1D array
If you begin with a 1D array like this one:
>>> arr = np . array ([ 1 , 2 , 3 , 4 , 5 , vi , vii , viii ]) Y'all tin can reverse it with:
>>> reversed_arr = np . flip ( arr ) If you want to impress your reversed array, y'all can run:
>>> print ( 'Reversed Array: ' , reversed_arr ) Reversed Array: [8 seven half dozen 5 iv 3 ii i] Reversing a 2D array
A 2D array works much the same way.
If yous outset with this assortment:
>>> arr_2d = np . array ([[ i , 2 , 3 , iv ], [ v , half-dozen , 7 , 8 ], [ ix , 10 , 11 , 12 ]]) Y'all can contrary the content in all of the rows and all of the columns with:
>>> reversed_arr = np . flip ( arr_2d ) >>> impress ( reversed_arr ) [[12 11 10 9] [ 8 7 vi 5] [ iv 3 two 1]] You can easily reverse just the rows with:
>>> reversed_arr_rows = np . flip ( arr_2d , axis = 0 ) >>> print ( reversed_arr_rows ) [[ 9 ten xi 12] [ 5 6 7 8] [ ane 2 three iv]] Or reverse just the columns with:
>>> reversed_arr_columns = np . flip ( arr_2d , axis = 1 ) >>> impress ( reversed_arr_columns ) [[ four 3 ii 1] [ 8 seven 6 5] [12 xi x 9]] You can also reverse the contents of only one cavalcade or row. For example, you can contrary the contents of the row at alphabetize position 1 (the second row):
>>> arr_2d [ 1 ] = np . flip ( arr_2d [ 1 ]) >>> impress ( arr_2d ) [[ 1 2 3 4] [ 8 vii half-dozen 5] [ 9 10 11 12]] You can too reverse the column at index position 1 (the second cavalcade):
>>> arr_2d [:, 1 ] = np . flip ( arr_2d [:, 1 ]) >>> print ( arr_2d ) [[ 1 10 3 4] [ eight seven half dozen v] [ 9 2 11 12]] Read more most reversing arrays at flip .
Reshaping and flattening multidimensional arrays¶
This section covers .flatten() , ravel()
There are ii popular ways to flatten an assortment: .flatten() and .ravel() . The main difference between the ii is that the new assortment created using ravel() is really a reference to the parent array (i.eastward., a "view"). This means that any changes to the new assortment will affect the parent array as well. Since ravel does not create a copy, it'due south memory efficient.
If y'all start with this array:
>>> x = np . array ([[ 1 , ii , 3 , iv ], [ 5 , half-dozen , seven , 8 ], [ nine , 10 , eleven , 12 ]]) Yous tin can apply flatten to flatten your array into a 1D assortment.
>>> 10 . flatten () array([ ane, 2, three, four, 5, 6, 7, 8, 9, 10, xi, 12]) When you use flatten , changes to your new array won't change the parent array.
For case:
>>> a1 = ten . flatten () >>> a1 [ 0 ] = 99 >>> print ( x ) # Original array [[ 1 2 iii iv] [ v six 7 8] [ nine ten 11 12]] >>> impress ( a1 ) # New array [99 2 3 iv 5 six 7 8 9 10 xi 12] But when yous use ravel , the changes y'all make to the new assortment will affect the parent assortment.
For instance:
>>> a2 = ten . ravel () >>> a2 [ 0 ] = 98 >>> print ( ten ) # Original assortment [[98 2 3 4] [ v 6 7 8] [ 9 10 11 12]] >>> print ( a2 ) # New array [98 two 3 4 5 6 7 8 ix 10 xi 12] Read more than most flatten at ndarray.flatten and ravel at ravel .
How to access the docstring for more data¶
This section covers help() , ? , ??
When information technology comes to the data science ecosystem, Python and NumPy are congenital with the user in listen. Ane of the all-time examples of this is the built-in access to documentation. Every object contains the reference to a string, which is known as the docstring. In nearly cases, this docstring contains a quick and concise summary of the object and how to use it. Python has a built-in assistance() role that tin help you access this information. This means that near any time y'all need more information, you can use help() to chop-chop find the data that you need.
For example:
>>> help ( max ) Assist on congenital-in role max in module builtins: max(...) max(iterable, *[, default=obj, key=func]) -> value max(arg1, arg2, *args, *[, key=func]) -> value With a single iterable statement, return its biggest detail. The default keyword-only statement specifies an object to return if the provided iterable is empty. With ii or more arguments, return the largest argument. Because access to additional information is and so useful, IPython uses the ? character equally a shorthand for accessing this documentation along with other relevant information. IPython is a control shell for interactive computing in multiple languages. You can find more than data nearly IPython here.
For example:
In [0]: max? max(iterable, *[, default=obj, cardinal=func]) -> value max(arg1, arg2, *args, *[, cardinal=func]) -> value With a single iterable argument, render its biggest item. The default keyword-only statement specifies an object to return if the provided iterable is empty. With two or more arguments, return the largest argument. Type: builtin_function_or_method You lot can even utilise this annotation for object methods and objects themselves.
Let's say you create this array:
>>> a = np . array ([ 1 , two , 3 , 4 , 5 , 6 ]) Then you tin can obtain a lot of useful information (starting time details nearly a itself, followed by the docstring of ndarray of which a is an instance):
In [1]: a? Type: ndarray String grade: [ane ii 3 iv 5 6] Length: 6 File: ~/anaconda3/lib/python3.7/site-packages/numpy/__init__.py Docstring: <no docstring> Class docstring: ndarray(shape, dtype=float, buffer=None, commencement=0, strides=None, club=None) An assortment object represents a multidimensional, homogeneous array of stock-still-size items. An associated information-type object describes the format of each element in the array (its byte-society, how many bytes information technology occupies in retentiveness, whether it is an integer, a floating point number, or something else, etc.) Arrays should exist synthetic using `assortment`, `zeros` or `empty` (refer to the Meet Also section below). The parameters given here refer to a low-level method (`ndarray(...)`) for instantiating an array. For more information, refer to the `numpy` module and examine the methods and attributes of an array. Parameters ---------- ( for the __new__ method ; see Notes below ) shape : tuple of ints Shape of created array . ... This besides works for functions and other objects that you create. Simply remember to include a docstring with your part using a cord literal ( """ """ or ''' ''' around your documentation).
For example, if you create this office:
>>> def double ( a ): ... '''Return a * two''' ... return a * two You lot tin obtain information nearly the function:
In [ii]: double? Signature: double(a) Docstring: Render a * two File: ~/Desktop/<ipython-input-23-b5adf20be596> Type: function You can achieve another level of data by reading the source code of the object y'all're interested in. Using a double question marker ( ?? ) allows you to access the source lawmaking.
For example:
In [3]: double?? Signature: double(a) Source: def double(a): '''Return a * 2''' render a * ii File: ~/Desktop/<ipython-input-23-b5adf20be596> Type: function If the object in question is compiled in a language other than Python, using ?? will return the same information as ? . You'll find this with a lot of built-in objects and types, for example:
In [4]: len? Signature: len(obj, /) Docstring: Return the number of items in a container. Blazon: builtin_function_or_method and :
In [5]: len?? Signature: len(obj, /) Docstring: Render the number of items in a container. Type: builtin_function_or_method have the same output because they were compiled in a programming language other than Python.
Working with mathematical formulas¶
The ease of implementing mathematical formulas that work on arrays is 1 of the things that brand NumPy so widely used in the scientific Python customs.
For case, this is the mean square error formula (a central formula used in supervised machine learning models that deal with regression):
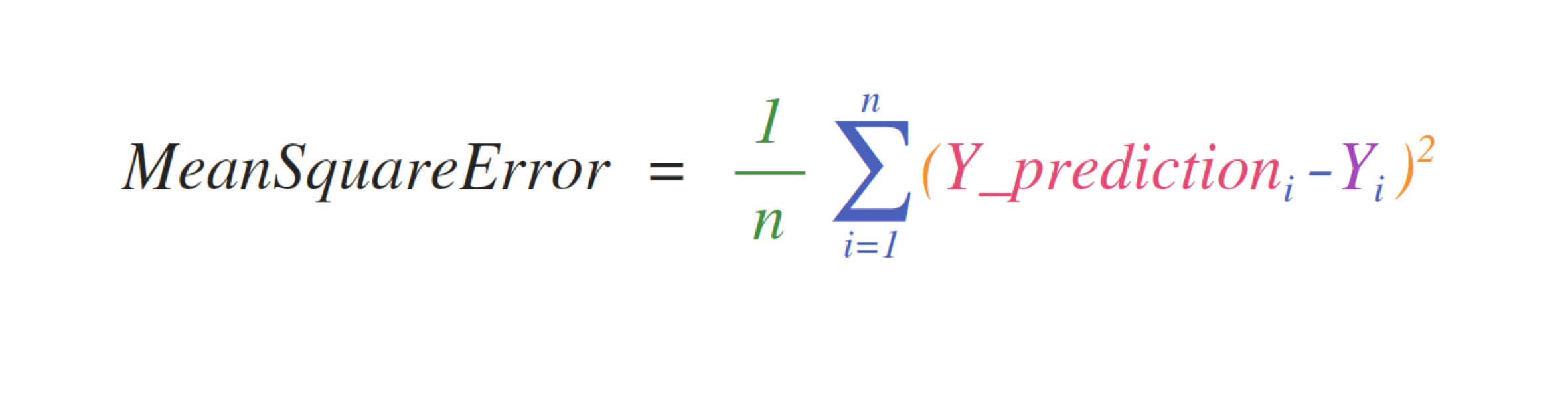
Implementing this formula is simple and straightforward in NumPy:
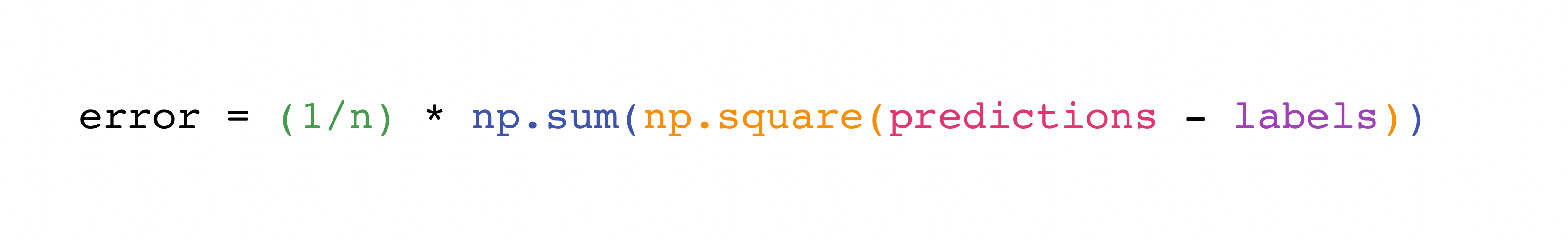
What makes this piece of work so well is that predictions and labels tin can comprise one or a g values. They but need to exist the same size.
You can visualize it this way:
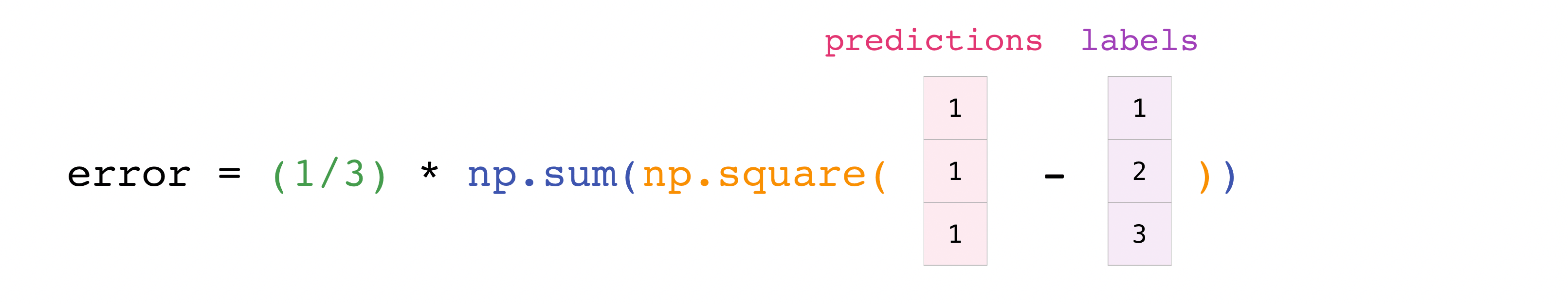

In this example, both the predictions and labels vectors contain three values, meaning n has a value of iii. After nosotros acquit out subtractions the values in the vector are squared. And so NumPy sums the values, and your result is the mistake value for that prediction and a score for the quality of the model.
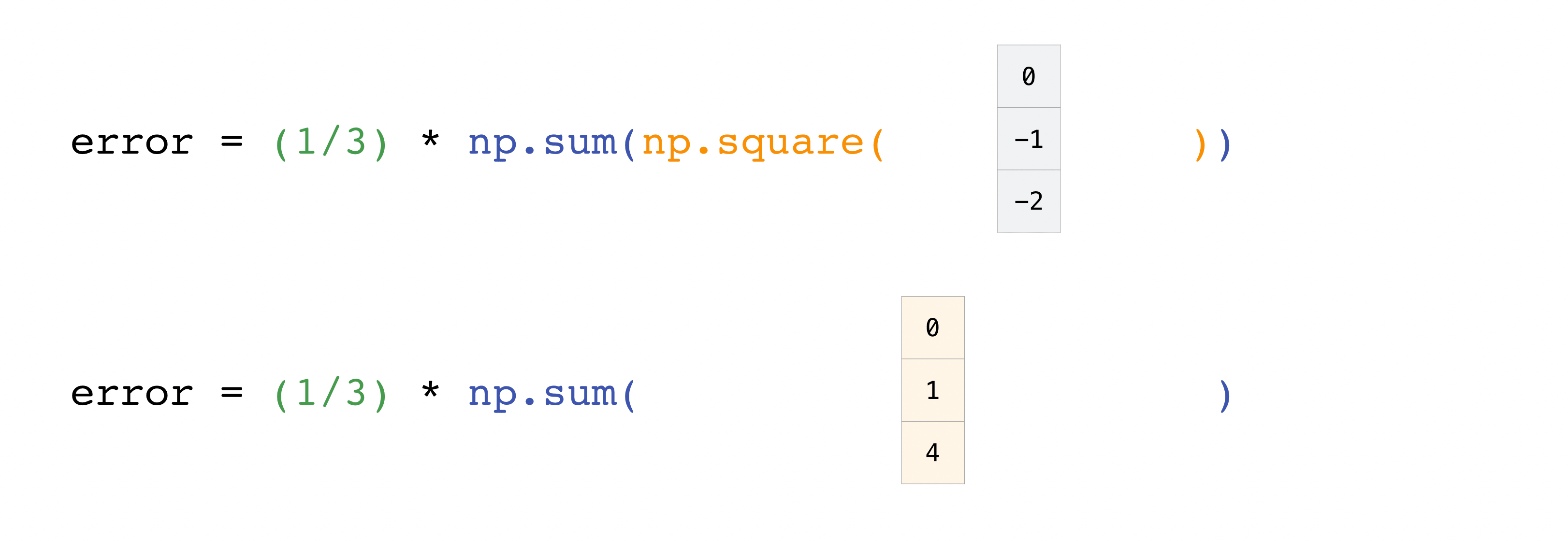

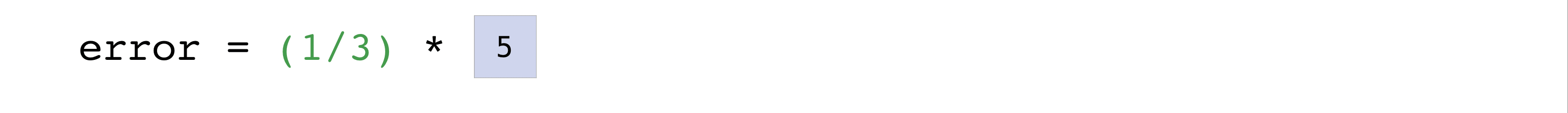
How to salvage and load NumPy objects¶
This section covers np.relieve , np.savez , np.savetxt , np.load , np.loadtxt
Y'all will, at some betoken, want to save your arrays to deejay and load them back without having to re-run the lawmaking. Fortunately, at that place are several ways to save and load objects with NumPy. The ndarray objects tin can be saved to and loaded from the disk files with loadtxt and savetxt functions that handle normal text files, load and save functions that handle NumPy binary files with a .npy file extension, and a savez function that handles NumPy files with a .npz file extension.
The .npy and .npz files store data, shape, dtype, and other information required to reconstruct the ndarray in a style that allows the array to be correctly retrieved, fifty-fifty when the file is on another machine with dissimilar architecture.
If you desire to shop a single ndarray object, shop it as a .npy file using np.salvage . If you want to store more than than i ndarray object in a unmarried file, relieve information technology as a .npz file using np.savez . You can also save several arrays into a single file in compressed npz format with savez_compressed .
Information technology's like shooting fish in a barrel to save and load and array with np.relieve() . Just make sure to specify the array you want to salve and a file name. For example, if you create this array:
>>> a = np . array ([ i , 2 , 3 , four , 5 , vi ]) You can salvage it as "filename.npy" with:
>>> np . save ( 'filename' , a ) You can use np.load() to reconstruct your array.
>>> b = np . load ( 'filename.npy' ) If y'all want to check your array, yous can run::
>>> print ( b ) [1 two iii 4 5 6] You can salve a NumPy array as a obviously text file like a .csv or .txt file with np.savetxt .
For example, if y'all create this array:
>>> csv_arr = np . array ([ 1 , 2 , 3 , iv , 5 , 6 , 7 , viii ]) You can hands save information technology as a .csv file with the proper name "new_file.csv" similar this:
>>> np . savetxt ( 'new_file.csv' , csv_arr ) You can rapidly and hands load your saved text file using loadtxt() :
>>> np . loadtxt ( 'new_file.csv' ) array([i., 2., 3., 4., 5., 6., 7., 8.]) The savetxt() and loadtxt() functions accept additional optional parameters such every bit header, footer, and delimiter. While text files tin can exist easier for sharing, .npy and .npz files are smaller and faster to read. If you need more sophisticated treatment of your text file (for example, if you lot need to work with lines that comprise missing values), you will want to use the genfromtxt office.
With savetxt , you lot can specify headers, footers, comments, and more.
Learn more than almost input and output routines here.
Importing and exporting a CSV¶
It'due south uncomplicated to read in a CSV that contains existing information. The best and easiest way to do this is to use Pandas.
>>> import pandas as pd >>> # If all of your columns are the same type: >>> 10 = pd . read_csv ( 'music.csv' , header = 0 ) . values >>> print ( ten ) [['Billie Vacation' 'Jazz' 1300000 27000000] ['Jimmie Hendrix' 'Rock' 2700000 70000000] ['Miles Davis' 'Jazz' 1500000 48000000] ['SIA' 'Pop' 2000000 74000000]] >>> # You lot can also just select the columns you demand: >>> ten = pd . read_csv ( 'music.csv' , usecols = [ 'Artist' , 'Plays' ]) . values >>> print ( x ) [['Billie Holiday' 27000000] ['Jimmie Hendrix' 70000000] ['Miles Davis' 48000000] ['SIA' 74000000]] 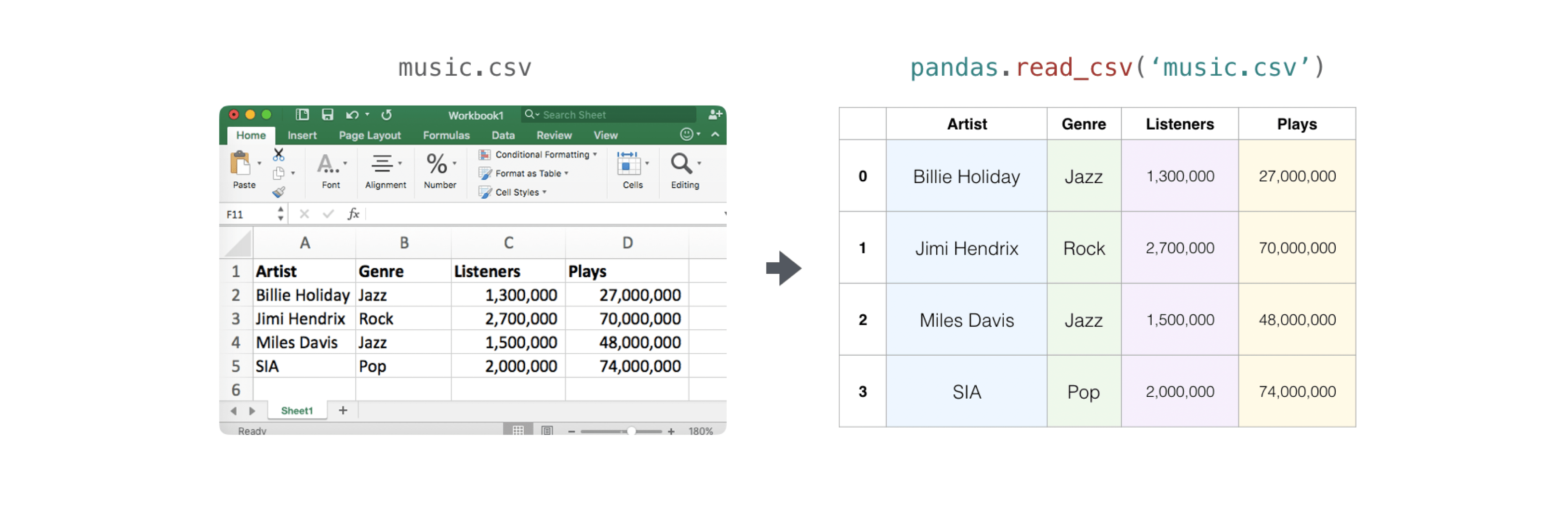
Information technology'south simple to utilise Pandas in order to export your array as well. If you are new to NumPy, yous may desire to create a Pandas dataframe from the values in your array and then write the data frame to a CSV file with Pandas.
If you created this array "a"
>>> a = np . array ([[ - 2.58289208 , 0.43014843 , - 1.24082018 , 1.59572603 ], ... [ 0.99027828 , 1.17150989 , 0.94125714 , - 0.14692469 ], ... [ 0.76989341 , 0.81299683 , - 0.95068423 , 0.11769564 ], ... [ 0.20484034 , 0.34784527 , 1.96979195 , 0.51992837 ]]) You could create a Pandas dataframe
>>> df = pd . DataFrame ( a ) >>> print ( df ) 0 1 2 three 0 -2.582892 0.430148 -1.240820 1.595726 1 0.990278 1.171510 0.941257 -0.146925 2 0.769893 0.812997 -0.950684 0.117696 3 0.204840 0.347845 1.969792 0.519928 You can easily relieve your dataframe with:
And read your CSV with:
>>> information = pd . read_csv ( 'pd.csv' ) 
Y'all can as well save your assortment with the NumPy savetxt method.
>>> np . savetxt ( 'np.csv' , a , fmt = ' %.2f ' , delimiter = ',' , header = '1, ii, three, 4' ) If y'all're using the command line, you can read your saved CSV any time with a command such every bit:
$ true cat np.csv # 1, 2, iii, 4 -2.58,0.43,-1.24,1.threescore 0.99,1.17,0.94,-0.15 0.77,0.81,-0.95,0.12 0.20,0.35,1.97,0.52 Or y'all can open the file whatever time with a text editor!
If you're interested in learning more about Pandas, take a expect at the official Pandas documentation. Learn how to install Pandas with the official Pandas installation information.
Plotting arrays with Matplotlib¶
If you need to generate a plot for your values, it'south very unproblematic with Matplotlib.
For example, yous may take an assortment like this i:
>>> a = np . array ([ two , 1 , 5 , 7 , 4 , 6 , eight , 14 , ten , 9 , 18 , 20 , 22 ]) If yous already have Matplotlib installed, y'all can import it with:
>>> import matplotlib.pyplot as plt # If you lot're using Jupyter Notebook, you may too want to run the following # line of code to display your code in the notebook: %matplotlib inline All you need to practice to plot your values is run:
>>> plt . plot ( a ) # If you are running from a command line, you may need to do this: # >>> plt.evidence() 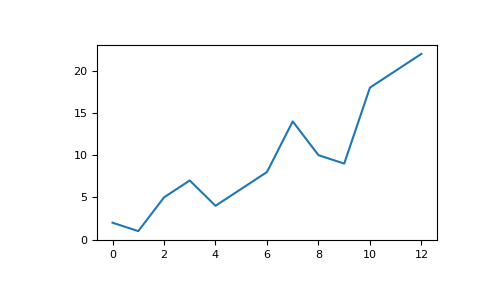
For example, you can plot a 1D array like this:
>>> x = np . linspace ( 0 , five , 20 ) >>> y = np . linspace ( 0 , 10 , twenty ) >>> plt . plot ( x , y , 'majestic' ) # line >>> plt . plot ( 10 , y , 'o' ) # dots 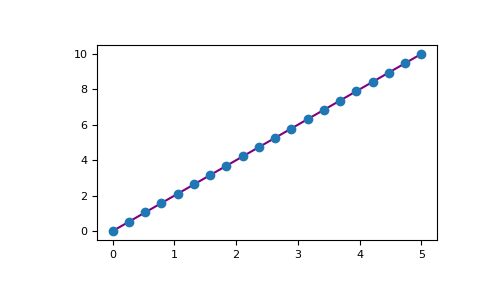
With Matplotlib, y'all take admission to an enormous number of visualization options.
>>> fig = plt . figure () >>> ax = fig . add_subplot ( projection = '3d' ) >>> X = np . arange ( - 5 , 5 , 0.15 ) >>> Y = np . arange ( - 5 , 5 , 0.15 ) >>> X , Y = np . meshgrid ( X , Y ) >>> R = np . sqrt ( Ten ** 2 + Y ** ii ) >>> Z = np . sin ( R ) >>> ax . plot_surface ( Ten , Y , Z , rstride = i , cstride = i , cmap = 'viridis' ) 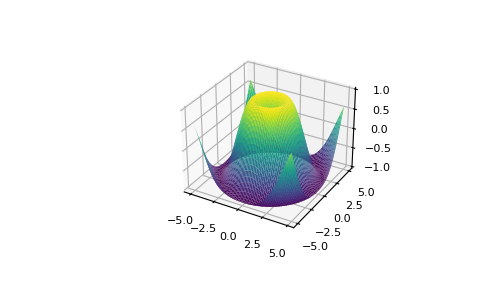
To read more about Matplotlib and what it tin practice, take a look at the official documentation. For directions regarding installing Matplotlib, see the official installation section.
Image credits: Jay Alammar http://jalammar.github.io/
Source: https://numpy.org/doc/stable/user/absolute_beginners.html
0 Response to "If We Want to Store an Array of Structures Into a Disk File, What File Type Should We Choose?"
Post a Comment